Informations générales
Date : le 23 novembre 2021 de 9h00 à 17h30
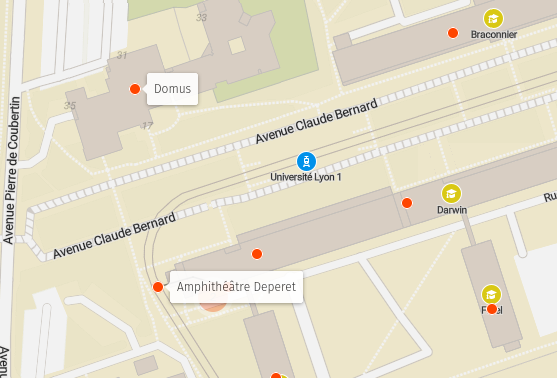
Lieu : Amphi Deperet, bâtiment Darwin, Campus de la Doua, Lyon - Accès
Organisateurs : Russ Harmer, Laurence Calzone, Cédric Lhoussaine, Loïc Paulevé et Élisabeth Remy
Cette sixième édition des journées annuelles du GT Bioss se déroulera juste avant la journée nationale du GDR BiM qui aura lieu le 24 novembre à l’Université Claude Bernard.
Inscription
L’inscription, gratuite mais obligatoire, se fait via la page du GDR BiM de l’évènement.
Programme
La journée se déroulera entre 9h et 17h30 dans l’amphi Deperet, bâtiment Darwin, Campus de la Doua, Lyon - Accès.
Orateurs invités:
- Alberto Valdeolivas (Roche Pharma Research and Early Development, Pharmaceutical Sciences, Roche Innovation Center Basel, Basel, Switzerland)
- Delphine Ropers (Inria Grenoble – Rhône-Alpes)
- Thomas Guillemaud et Celine Scornavacca (Peer Community In - PCI)
Déroulé de la journée
Amphi Deperet, bâtiment Darwin
9h00 - 9h25 Accueil café
9h30 - 9h45 - Présentation du GT Bioss
9h45 - 10h30 - Exposé invité : Alberto Valdeolivas - Exploring colorectal cancer heterogeneity with Spatial Transcriptomics
10h30 - 11h50 - Exposés courts
- 10h30 - 10h50: Déborah Boyenval - Mammalian cell cycle: formalizing phases
- 10h50 - 11h10: Anthony Baptista - Universal Multilayer Network Exploration by Random Walk with Restart
- 11h10 - 11h30: Arnaud Belcour - Estimating the metabolic capacity of a community from taxonomic assignation
- 11h30 - 11h50: Gautier Stoll - Modeling Immunogenic Cell Death with a stochastic Boolean approach
Salle VIP du restaurant DOMUS (bâtiment en face de Darwin)
12h00 - 13h25 Repas
Amphi Deperet, bâtiment Darwin
13h30 - 14h15 - Exposé invité : Delphine Ropers - Model-based interpretation of high-throughput data: Integrative analysis of bacterial mRNA decay
14h15 - 15h35 - Exposés courts
- 14h15 - 14h35 Honglu Sun - Learning and stability analysis of a hybrid modeling of dynamical biological systems
- 14h35 - 14h55 Albin Salazar - Interval-based coarse graining of a reaction network
- 14h55 - 15h15 Matthieu Bougueon - A kappa model for hepatic stellate cells activation by TGFB1
- 15h15 - 15h35 Nicolas Levy - ENSnano: a software for designing DNA 3D nanostructure
15h40 - 16h10 - Pause café
16h15 - 17h15 - Exposé + Table ronde PCI - Thomas Guillemaud et Celine Scornavacca
Résumés
Alberto Valdeolivas - Exploring colorectal cancer heterogeneity with Spatial Transcriptomics.
Colorectal cancer (CRC) is a leading cause of cancer-related death worldwide with over 1.85 million diagnosed cases and 850 000 deaths annually. CRC mortality rates have improved in recent years as a result of treatments tailored to the molecular and pathological features of the different groups of patients. The integrative analysis of large scale DNA and RNA sequencing data profoundly contributed to characterize CRC diversity and resulted in the classification of CRC into four consensus molecular subtypes (CMS) with distinguishing features. However, the spatial heterogeneity of CMS subtypes and its potential relevance for treatment decisions remains elusive.
In the last decade, we have witnessed substantial advances in next generation sequencing and multiplex bioimaging based technologies. The effective combination of these approaches paved the development of Spatial Transcriptomics (ST). ST enables measuring of gene expression levels throughout sample space, preserving the cellular distribution in the tissue under analysis. These expression maps are crucial for understanding the influence of cellular and tissue organization in their biological function, and can therefore lead to new insights in a large variety of areas, such as disease understanding.
In this study, we applied ST and a deconvolution-based approach to decipher the CMS and spatial heterogeneity of seven CRC samples. We further used our ST data to investigate cell communication at the tumor stroma interface. Our results suggest large variability among CMS2 tumors at the gene expression, pathway and transcription factor activity levels. In addition, we found substantial differences in the molecular features of their surrounding stroma.
Delphine Ropers - Model-based interpretation of high-throughput data: Integrative analysis of bacterial mRNA decay
Intracellular levels of bacterial mRNAs are tightly controlled, with myriads of regulatory factors adjusting transcription and decay rates to environment. Contrary to transcription, little is known about how cells coordinate the changes of mRNA stability. Temporal transcriptomics data provide unique opportunities to address this question, but their mechanistic interpretation remains challenging. We develop an approach based on the mechanistic modelling of bacterial mRNA degradation at the whole-cell level. We estimate the distribution of physiologically-interpretable model parameters using non-linear mixed effect modelling and time-series omics data sets. Results reveal a new global regulatory mechanism of mRNA decay, competition between mRNAs for binding to the degradation machinery. It accounts for the massive adjustment of mRNA stabilities when growth conditions change. Sole 15% of mRNAs are additionally regulated by specific factors such as small RNAs and ribonucleoproteins. Our approach is able to cope with noisy omics data. It is modular and scalable to more complex models and data.
Déborah Boyenval - Mammalian cell cycle: formalizing phases
Checkpoints ensure the integrity of DNA during the cell cycle, which is a succession of molecular and cellular events leading to the division of a mother cell into two genetically identical daughter cells. The DNA is first duplicated (S-phase), then equally distributed between the opposing poles of the cell, which finally divides into two independent daughter cells (M-phase). Two additional phases depict the process of cell preparation before S and M phases: respectively the G1 and G2 phases. The checkpoint concept is fundamentally discrete, described as follows: a checkpoint prevents any event that initiates a phase from taking place before the completion of all the events of the previous phase. Many cell cycle models investigating checkpoints exist but, so far to our knowledge, very few qualitative models have yet attempted to formalize the discrete concept of checkpoint itself, notably because the notion of cell cycle phase is still fuzzy. Therefore, we propose a qualitative modeling study of cell cycle regulation dedicated to the logical specification of the G1, S, G2 and M phases, and the G1/S, S/G2, G2/M and mitosis exit checkpoints. This study has been made possible by using two types of formal methods: the “genetically modified Hoare logic” [1] and the model-checking for CTL [2]. The TotemBioNet tool efficiently combines these two methods to exhaustively identify the parameterizations (which govern the dynamics of the regulatory graph) compatible with all formalized biological knowledge [3]. Starting from a qualitative model of cell cycle progression regulation (Behaegel et al. [4]), the cell cycle was defined by a Hoare triple of the form {precondition}path{postcondition}, where the precondition is a single initial state, the path is a sequence of discrete events, and the postcondition is the single final state. The path was then divided into four canonical phases in order to identify non-permutable key events, which will constitute the main rule of the predicate checkpoint(phasei, phasei+1). Since the order of events within a phase is not necessarily known, the predicate (implemented in Prolog) includes rules which call TotemBioNet to extract all the orders of events of a phase compatible with all the biological knowledge stored within a cell cycle model.
[1] Gilles Bernot, Jean-Paul Comet, Zohra Khalis, Adrien Richard, and Olivier Roux. A genetically modified hoare logic. Theoretical Computer Science, 765, 06 2015.
[2] G. Bernot, J.-P. Comet, A. Richard, and J. Guespin. Application of formal methods to biological regulatory networks: Extending Thomas’ asynchronous logical approach with temporal logic. Journal of TheoreticalBiology, 229(3):339–347, 2004.
[3] Déborah Boyenval, Gilles Bernot, Hélène Collavizza, and Jean-Paul Comet. What is a cell cycle checkpoint? the totembionet answer. In 18th International Conference on Computational Methods in Systems Biology (CMSB 2020), 2020.
[4] J. Behaegel, J.-P. Comet, G. Bernot, E. Cornillon, and F. Delaunay. A hybrid model of cell cycle in mammals. J. Bioinformatics and Comput. Biol., 14(1):1640001 [17 pp.], 2016.
Anthony Baptista - Universal Multilayer Network Exploration by Random Walk with Restart
The amount and variety of data are increasing drastically for several years. These data are often represented as networks, which are then explored with approaches arising from network theory. Recent years have witnessed the extension of network exploration methods to leverage more complex and richer network frameworks. Random walks, for instance, have been extended to explore multilayer networks. However, current random walk approaches are limited in the combination and heterogeneity of network layers they can handle. New analytical and numerical random walk methods are needed to cope with the increasing diversity and complexity of multilayer networks.
I present here MultiXrank, a Python package that enables Random Walk with Restart (RWR) on any kind of multilayer network with an optimized implementation. This package is supported by a universal mathematical formulation of the RWR. I evaluated MultiXrank with leave-one-out cross-validation and link prediction, and introduced protocols to measure the impact of the addition or removal of multilayer network data on prediction performances. Furthermore, I measured the sensitivity of MultiXrank to input parameters by in-depth exploration of the parameter space. Finally, I will illustrate the versatility of MultiXrank with different use-cases of unsupervised node prioritization and supervised classification in the context of human genetic diseases.
Arnaud Belcour - Estimating the metabolic capacity of a community from taxonomic assignation
In this talk, I will present two methods for elucidating which organisms in a microbial community are associated with a specific metabolic function. Microbial community samples can be described by sequencing gene markers (such as the 16S rRNA gene), which are assigned to specific taxon using gene-sequence alignment methods. Although some functional profiles can be associated with these taxa, there is a lack of method for estimating the metabolic capacity of a community from the taxonomic assignations of a metagenomics sample, preventing the study of the metabolic dynamics of microbial communities.
The first method I developed estimates which protein are likely to be associated with a taxonomic assignation from gene markers, using queries on the Uniprot database. Once these proteins have been extracted their functional annotations are used to produce a draft metabolic network for the taxon. The second method, Metage2Metabo, aims at finding the key species that collaborate within the community with respect to the realization of an expected phenotype. Using constraint programming, this method processes large-scale datasets of microbial or taxon metabolic networks to highlight putative exchanges within the community and calculate the organisms involved in the production of metabolites of interest.
These two methods were combined to analyze microbial community data related to a biogas plant. By combining database interrogation, system biology, and sequence analysis, I will illustrate how our methods allow to find key community members associated with specific steps of the methanogenesis.
Gautier Stoll - Modeling Immunogenic Cell Death with a stochastic Boolean approach
The immunological cell death is a dying process that activates the immune system. It has been observed in tumor treatment by cytotoxic agent. It is a possible explanation of the efficacy of some chemotherapies.
This type of cell death has been intensively studied within mouse model. A complicated molecular mechanism emerges from these experimental studies, implying different pathways within different cell types, around a global positive feedback loop.
As a part of an ITMO project, we construct different version of mathematical models of Immunogenic Cell Death. For that, we used a stochastic Boolean approach within a software tool, UPMaBoSS, an extension of MaBoSS for dynamical modelling of heterogenous cell populations, integrating cell division, cell death and ligand-receptor interaction. MaBoSS itself is a continuous time Boolean modelling approach for pathway modeling.
We will present the tool UPMaBoSS. We will show the first results of our models: sensitive analysis and drug target prediction. Then, we will present to ongoing works: an extended Immunogenic Cell Death model and an extension of UPMaBoSS (called PopMaBoSS) that is able to tackle the individual variation of Immunogenic Cell Death induction.
Honglu Sun - Learning and stability analysis of a hybrid modeling of dynamical biological systems
Using mathematical models to study the dynamics of gene regulatory system is very important because of the complex nature of biological systems. The two most used formalisms are discrete models (like Boolean networks) and continuous models (like differential equations). The dynamics of discrete models are easy to analyze but sometimes not precise enough (for example, it is hard to identify damped oscillation in discrete models). Continuous models are more precise but their dynamics are sometimes hard to analyze. In past years, there are also some hybrid models which have been proposed as a way to build a bridge between approaches: they can be seen as a simplification of the continuous models or an extension of the discrete model. These hybrid models contain both continuous and discrete components. Our work is based on a class of hybrid models [1][2] which is an extension of discrete model. In this class of hybrid models, the state space is separated into several discrete states like discrete model and in each discrete state the temporal derivative of system is a constant vector so it can also make a continuous evolution of the system over time like differential equations. The objective of our work is to create a framework which firstly learns automatically a hybrid model of gene regulatory networks from biological data and then analyzes automatically the dynamical properties of the model. This work concerns two problems associated to the hybrid model: model learning (automatic identification of parameters) and analysis of dynamical properties. For the problem of model learning, we propose a learning method based on genetic algorithm which can directly learns models from time series data and shows the effectiveness of the method based on time series data generated by differential equations of several circadian clock systems. For the problem of dynamical properties analysis, we are working on formal methods to search for closed orbits and analyze the stability of the closed orbits based on Poincaré map. In future works, we would like to use our dynamical analysis methods to study the control problems of gene regulatory networks and exhibit the merits of our learning methods by applying them to a proper biological case study.
[1] Cornillon, E., Comet, J. P., Bernot, G., & Enée, G. (2016). Hybrid gene networks: a new framework and a software environment. advances in Systems and Synthetic Biology.
[2] Behaegel, J., Comet, J. P., & Folschette, M. (2017). Constraint identification using modified Hoare logic on hybrid models of gene networks. In 24th International Symposium on Temporal Representation and Reasoning (TIME 2017). Schloss Dagstuhl-Leibniz-Zentrum fuer Informatik.
Albin Salazar - Interval-based coarse graining of a reaction network
The combinatorial problem is a common feature that arises from the structure of mechanistic models in biology and (continuously) poses a challenge when developing large-scale descriptions of biological systems. For example, the analysis or inference of biological behaviours generated from Boolean network models with a couple of dozen variables becomes intractable. Thus, research efforts from various disciplines collaborate to reduce the space of possible solutions that pertain to, for instance, control targets in drug development, modulators of metabolic pathways, or key events in signalling pathways responsible for cell integrity. Consequently, we share a common goal: to capture such biological behaviours at a low level of representation. Generally, the focus of our work is to develop faithful model reduction techniques by constructing formal (modelling) tools. This is done by using methods from the field of Abstract Interpretation. As a case study, the context of these tools is to address problems of concentration- and time-scale separation in qualitative models of biological systems. As such, we show that techniques from Abstract Interpretation is capable of developing coarse-grained models of a probabilistic transition system. Additionally, we discuss how these models can be refined to capture the (stochastic) dynamics of multi-dimensional systems.
Matthieu Bougueon - A kappa model for hepatic stellate cells activation by TGFB1
All chronic hepatitis are associated with the development of fibrosis, which results in abnormal deposition of the extracellular matrix (ECM) leading to severe liver dysfunction. Fibrosis final stage, called cirrhosis, is the main risk of development of hepatocellular carcinoma (HCC). At the cellular level, hepatic stellate cells (HSCs) are major actors of fibrosis and tumor progression. Upon liver injury, HSCs are activated to repair tissue and are subsequently eliminated through three mechanisms: apoptosis, senescence and reversion, leading to a return to healthy status [5]. However, when the injury persists, HSCs remain activated with a myobroblastic phenotype, and extracellular matrix accumulates, leading to fibrosis, cirrhosis and cancer. Understanding the dynamics of HSC activation and their regulation by TGFB1 is essential to identify markers and therapeutic targets that may favor the resolution of fibrosis at the expense of its progression. For this purpose, we are developing a modelling approach using the Kappa language. Kappa is a rule-based language used for the rewriting of site graphs [1, 2, 4, 3] aiming at describing networks of interactions between occurrences of components, using a syntax inspired by chemistry. In this model, the components are occurrences of HSC in different states, and occurrences of the TGFB1 protein. Our preliminary results suggest a high plasticity of the HSC response to TGFB1 stimulation. Future work will focus on the integration of the ECM component networks that regulate TGFB1 availability.
[1] O. Andrei and H. Kirchner. A rewriting calculus for multigraphs with ports. Electr. Notes Theor. Comput. Sci., 219:67–82, 2008.
[2] V. Danos and C. Laneve. Formal molecular biology. Theoretical Computer Science, 325(1):69 – 110, 2004. Computational Systems Biology.
[3] A. Ehrlich, D. Duche, G. Ouedraogo, and Y. Nahmias. Challenges and opportunities in the design of liver-on-chip microdevices. Annual Review of Biomedical Engineering, 21(1):219–239, 2019. PMID: 31167098.
[4] J. R. Faeder, M. L. Blinov, B. Goldstein, and W. S. Hlavacek. Rule-based modeling of biochemical networks. Complexity, 10(4):22–41, 2005.
[5] T. Kisseleva and D. Brenner. Molecular and cellular mechanisms of liver fibrosis and its regression. Nature Reviews Gastroenterology & Hepatology, pages 1–16, 2020
Nicolas Levy - ENSnano: a software for designing DNA 3D nanostructure
Since the 1990s, increasingly complex nanostructures have been reliably obtained out of self-assembled DNA strands: from “simple” 2D shapes to 3D gears and articulated nano-objects, and even computing structures. The success of the assembly of these structures relies on a fine tuning of their structure to match the peculiar geometry of DNA helices. Various softwares have been developed to help the designer. These softwares provide various tools to visualize and edit the designed structure.
Our goal is to identify and develop abstractions for working with DNA nanostructures. The whole stake is to define a mental model that is both easy to apprehend and expressive enough to design complex structures. We then want to develop effective user interfaces to interact with that model. In this talk, we present a model that we develop and explain why we think it is an effective way to work with DNA. We also present ENSnano, a software for designing 3D DNA nanostructures that implements our model. ENSnano takes into account the latest discoveries regarding the geometry of the DNA double helix. The software is validated experimentally by wet-lab experiments whose results are taken into account for improving the software
Dernière modification le 23/11/2021